Running example
Running mitEA is as easy as running GOrilla (that is, very easy...). Just follow the example below:
Step 1: Choose "Homo sapiens"
Step 2: Paste the contents of this file to the textbox.
This example is the list
of down regulated proteins in HeLa cells following miR-1 transfection.
The data is from Sellbach et al, Nature 2008..
Step 3: Choose the target prediction tool - "TargetScan"
You can adjust the sensitivity of results using the "P-value threshold". Change threshold to 10^-5.
Click on "Search enriched miRNA targets"
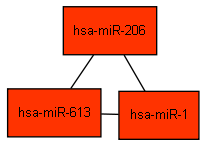
That's it. The following figure should appear on the screen:
